The Replication Network
Furthering the Practice of Replication in Economics
IN THE NEWS: Mother Jones (September 25, 2018)
[From the article, “This Cornell Food Researcher Has Had 13 Papers Retracted. How Were They Published in the First Place?” by Kiera Butler, published in Mother Jones]
“In 2015, I wrote a profile of Brian Wansink, a Cornell University behavioral science researcher who seemed to have it all: a high-profile lab at an elite university, more than 200 scientific studies to his name, a high-up government appointment, and a best-selling book.”
“…In January 2017, a team of researchers reviewed four of [Wansink’s] published papers and turned up 150 inconsistencies. Since then, in a slowly unfolding scandal, Wansink’s data, methods, and integrity have been publicly called into question. Last week, the Journal of the American Medical Association (JAMA) retracted six articles he co-authored. To date, a whopping 13 Wansink studies have been retracted.”
“… when I first learned of the criticisms of his work, I chalked it up to academic infighting and expected the storm to blow over. But as the scandal snowballed, the seriousness of the problems grew impossible to ignore. I began to feel foolish for having called attention to science that, however fun and interesting, has turned out to be so thin. Were there warning signs I missed? Maybe. But I wasn’t alone. Wansink’s work has been featured in countless major news outlets—the New York Times has called it “brilliantly mischievous.” And when Wansink was named head of the USDA in 2007, the popular nutrition writer Marion Nestle deemed it a “brilliant appointment.””
“Scientists bought it as well. Wansink’s studies made it through peer review hundreds of times—often at journals that are considered some of the most prestigious and rigorous in their fields. The federal government didn’t look too closely, either: The USDA based its 2010 dietary guidelines, in part, on Wansink’s work. So how did this happen?”
To read more, click here.
How Many Biases? Let Us Count the Ways
[From the article “Congratulations. Your Study Went Nowhere” by Aaron Carroll, published at http://www.nytimes.com]
“When we think of biases in research, the one that most often makes the news is a researcher’s financial conflict of interest. But another bias, one possibly even more pernicious, is how research is published and used in supporting future work.”
“A recent study in Psychological Medicine examined how four of these types of biases came into play in research on antidepressants.”
“… Publication bias refers to the decision on whether to publish results based on the outcomes found. “
“… Outcome reporting bias refers to writing up only the results in a trial that appear positive, while failing to report those that appear negative.”
“… Spin refers to using language, often in the abstract or summary of the study, to make negative results appear positive.”
“… Research becomes amplified by citation in future papers. The more it’s discussed, the more it’s disseminated both in future work and in practice. Positive studies were cited three times more than negative studies. This is citation bias.”
To read more, click here.
18,000 Retractions?
[From the video, “The Retraction Watch Database” by Ivan Oransky, posted at YouTube].
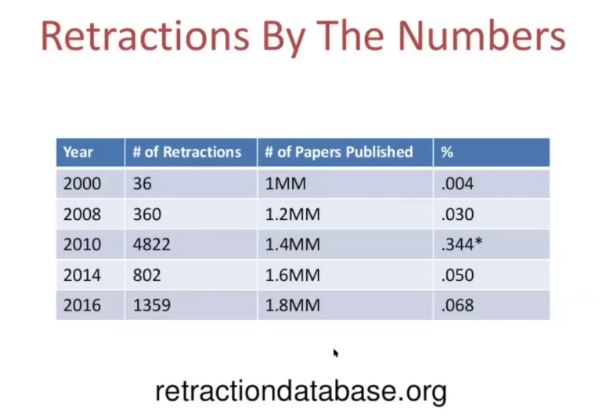
Ivan Oransky, MD, co-founder of the website Retraction Watch gave a talk at the Joint Roadmap for Open Science Tools Workshop at Berkeley in August. In this short video, Oranksy talks about how retracted papers continue to be cited after they have been retracted. 40% of the time when a retracted paper has been cited, there is no acknowledgment that the paper has been retracted (likely because the citing author did not know). And then there’s this:
To watch the full video (it’s only about 8 and a half minutes long), click here.
M Is For Pizza
[From the blog ““Tweeking”: The big problem is not where you think it is” by Andrew Gelman, posted at Statistical Modeling, Causal Inference, and Social Science]
“In her recent article about pizzagate, Stephanie Lee included this hilarious email from Brian Wansink, the self-styled “world-renowned eating behavior expert for over 25 years”:
“OK, what grabs your attention is that last bit about “tweeking” the data to manipulate the p-value, where Wansink is proposing research misconduct (from NIH: “Falsification: Manipulating research materials, equipment, or processes, or changing or omitting data or results such that the research is not accurately represented in the research record”).”
“But I want to focus on a different bit: “. . . although the stickers increase apple selection by 71% . . .””
“This is the type M (magnitude) error problem—familiar now to us, but not so familiar a few years ago to Brian Wansink, James Heckman, and other prolific researchers.”
To read more, click here.
IN THE NEWS: Bloomberg (September 18, 2018)
[From the article “Why Economics Is Having a Replication Crisis” by Noah Smith, published at http://www.bloomberg.com]
“By now, most people have heard of the replication crisis in psychology. When researchers try to recreate the experiments that led to published findings, only slightly more than half of the results tend to turn out the same as before. Biology and medicine are probably riddled with similar issues.”
“But what about economics? Experimental econ is akin to psychology, and has similar issues. But most of the economics research you read about doesn’t involve experiments — it’s empirical, meaning it relies on gathering data from the real world and analyzing it statistically. Statistical calculations suggest that there are probably a lot of unreliable empirical results getting published and publicized.”
“…That doesn’t mean that single results aren’t worth reporting or taking into account, but a single finding shouldn’t be enough to generate certainty about how the world works. In a universe filled with uncertainty, social science can’t progress by leaps and bounds — it must crawl forward, feeling its way inch by inch toward a little more truth.”
To read more, click here.





You must be logged in to post a comment.